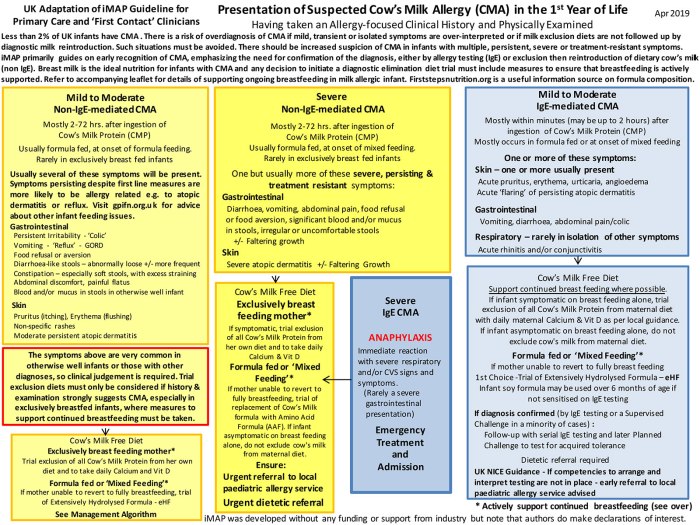
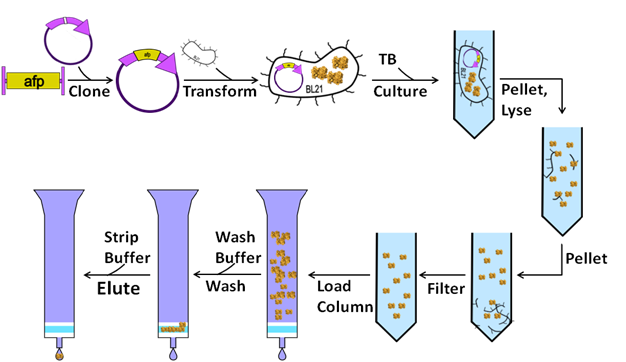
Immobilized Metal Affinity Chromatography (IMAC) for Metalloproteomics and Phosphoproteomics | Semantic Scholar
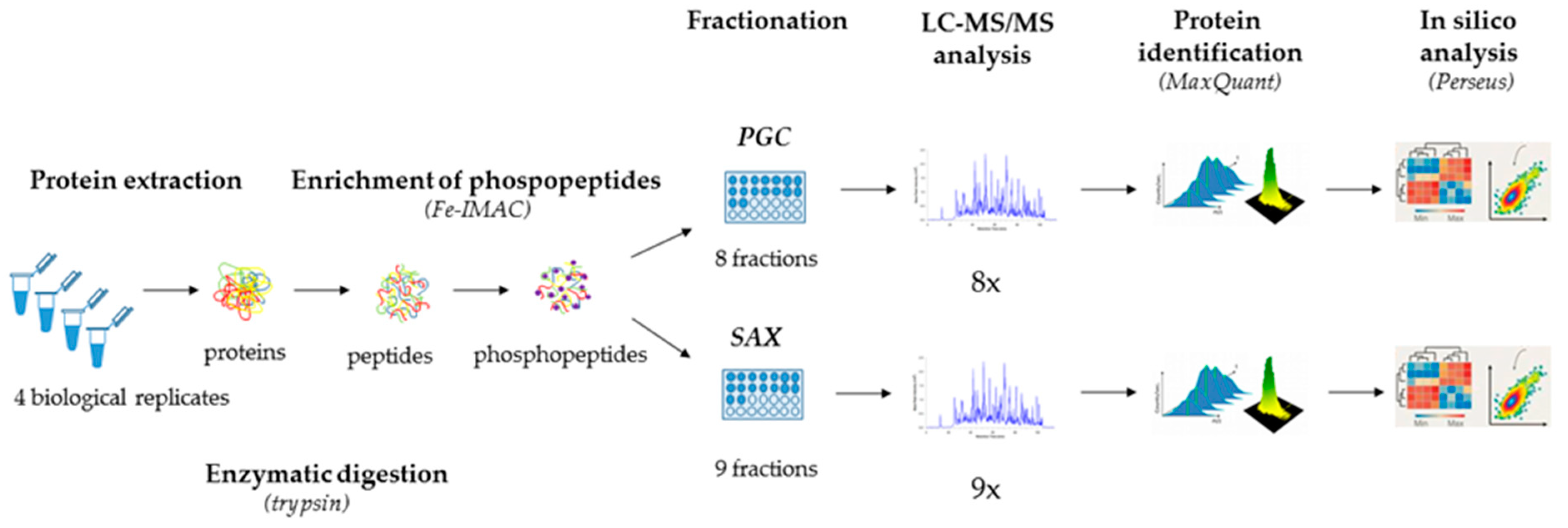
IJMS | Free Full-Text | Label-Free Quantitative Phosphoproteomics of the Fission Yeast Schizosaccharomyces pombe Using Strong Anion Exchange- and Porous Graphitic Carbon-Based Fractionation Strategies
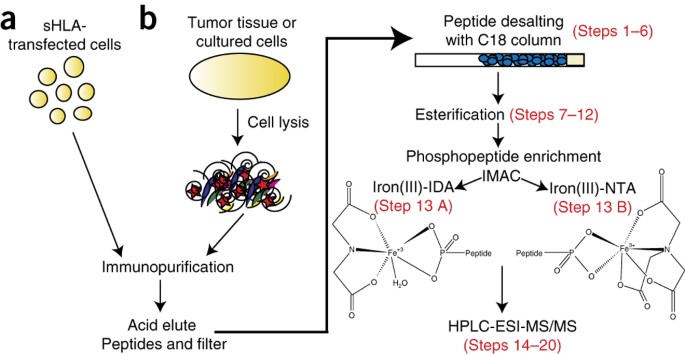
Complementary IMAC enrichment methods for HLA-associated phosphopeptide identification by mass spectrometry | Nature Protocols

ReSyn Biosciences on X: "In case you missed it, a fully automated STAR protocol incorporating our Ti-IMAC for "Unraveling virus-induced cellular signaling cascades by label-free quantitative phosphoproteomics" #Ti_IMAC https://t.co/hFQD9oQMYe https://t ...

Optimized IMAC−IMAC Protocol for Phosphopeptide Recovery from Complex Biological Samples | Journal of Proteome Research

Size exclusion chromatography of IMAC purified Nanobodies 1174, 1175 &... | Download Scientific Diagram

Optimized IMAC−IMAC Protocol for Phosphopeptide Recovery from Complex Biological Samples | Journal of Proteome Research
IMAC: An Interference-Aware Duty-Cycle MAC Protocol for Wireless Sensor Networks Employing Multipath Routing
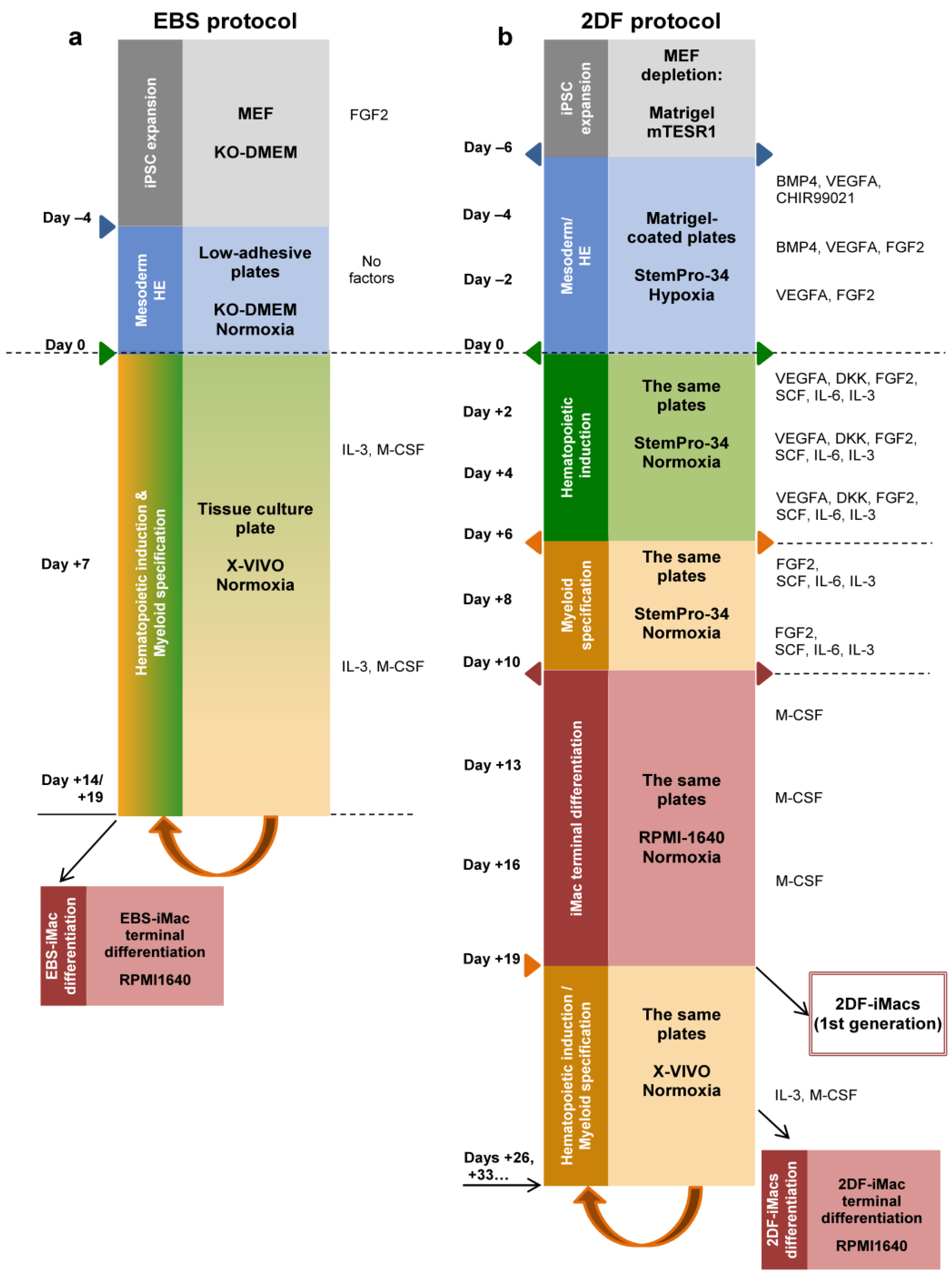
IJMS | Free Full-Text | iPSC-Derived Macrophages: The Differentiation Protocol Affects Cell Immune Characteristics and Differentiation Trajectories

USB C 3.2 Gen2 MacBook SSD Enclosure for Apple Flash SSDs 12+16 PIN MacBook Pro MacBook Air Mac Pro iMac from 2013 to 2017, with Adapter to Support M.2 NVMe SSDs (PCIe
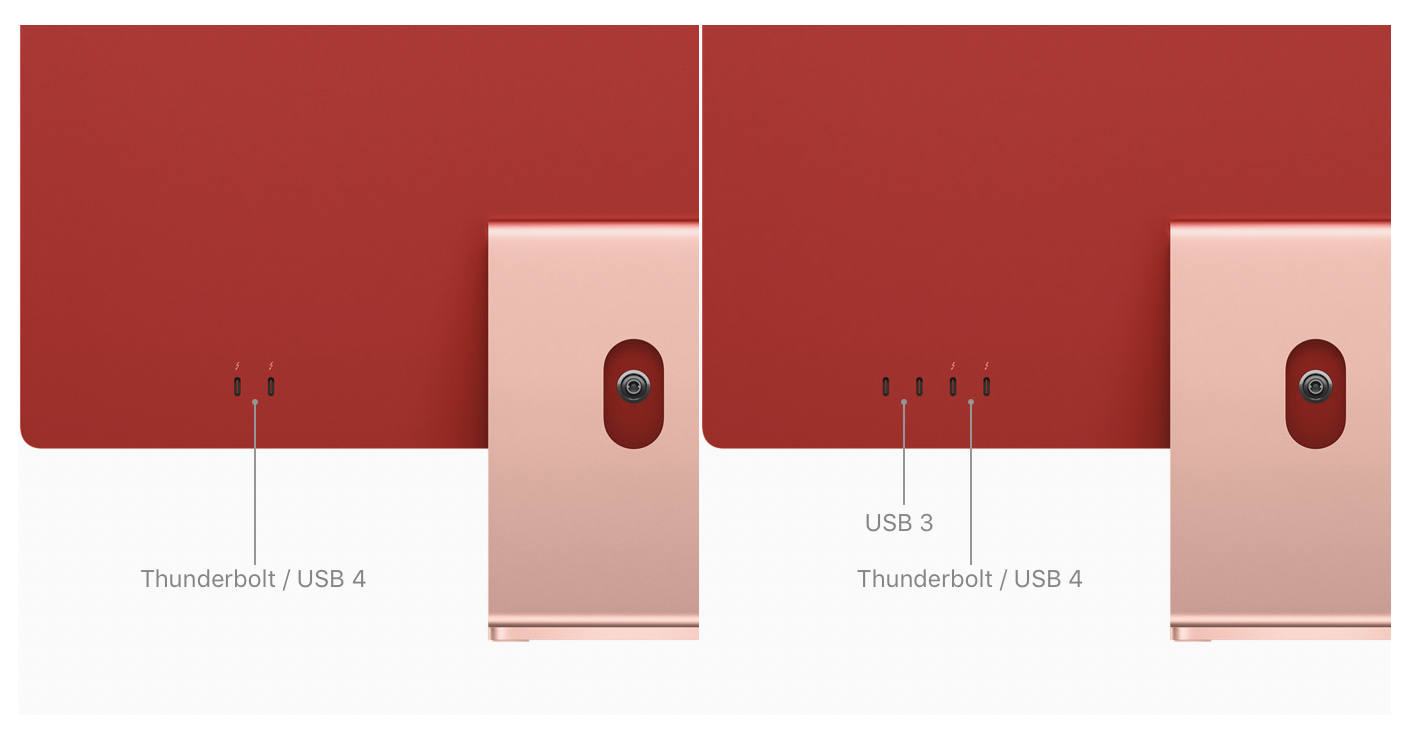
USB-C and Thunderbolt: Understanding Ports and Cables for Macs, iPhones, and iPads - The Mac Security Blog

Improving the yield of recalcitrant Nanobodies® by simple modifications to the standard protocol - ScienceDirect

Immobilized Metal Affinity Chromatography (IMAC) for Metalloproteomics and Phosphoproteomics | Semantic Scholar